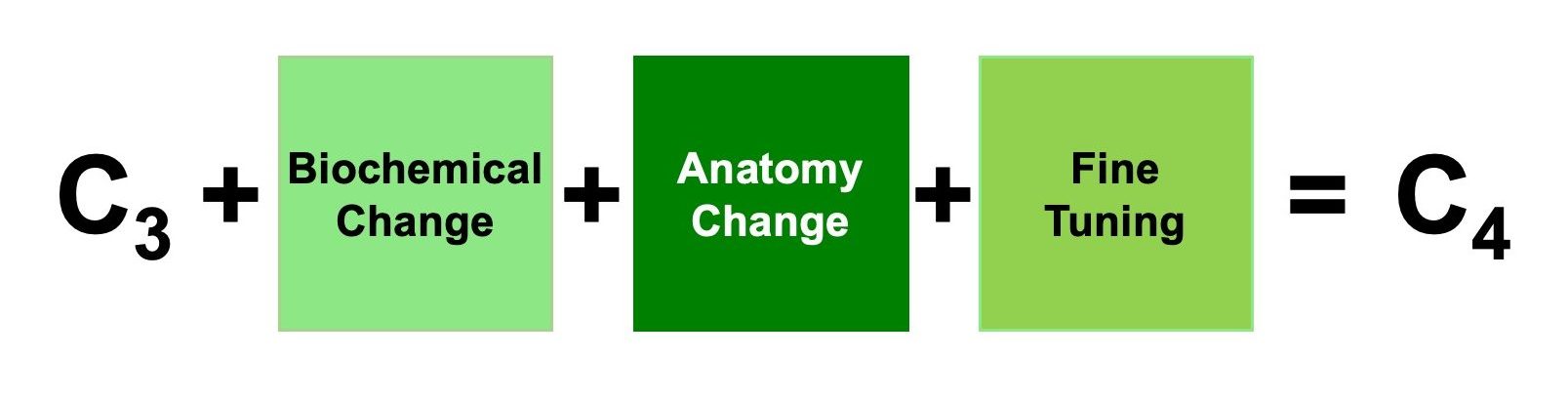
Engineering the C4 pathway into a C3 plant requires manipulation of both anatomical and biochemical traits and our project is organized into two workstreams around these objectives.
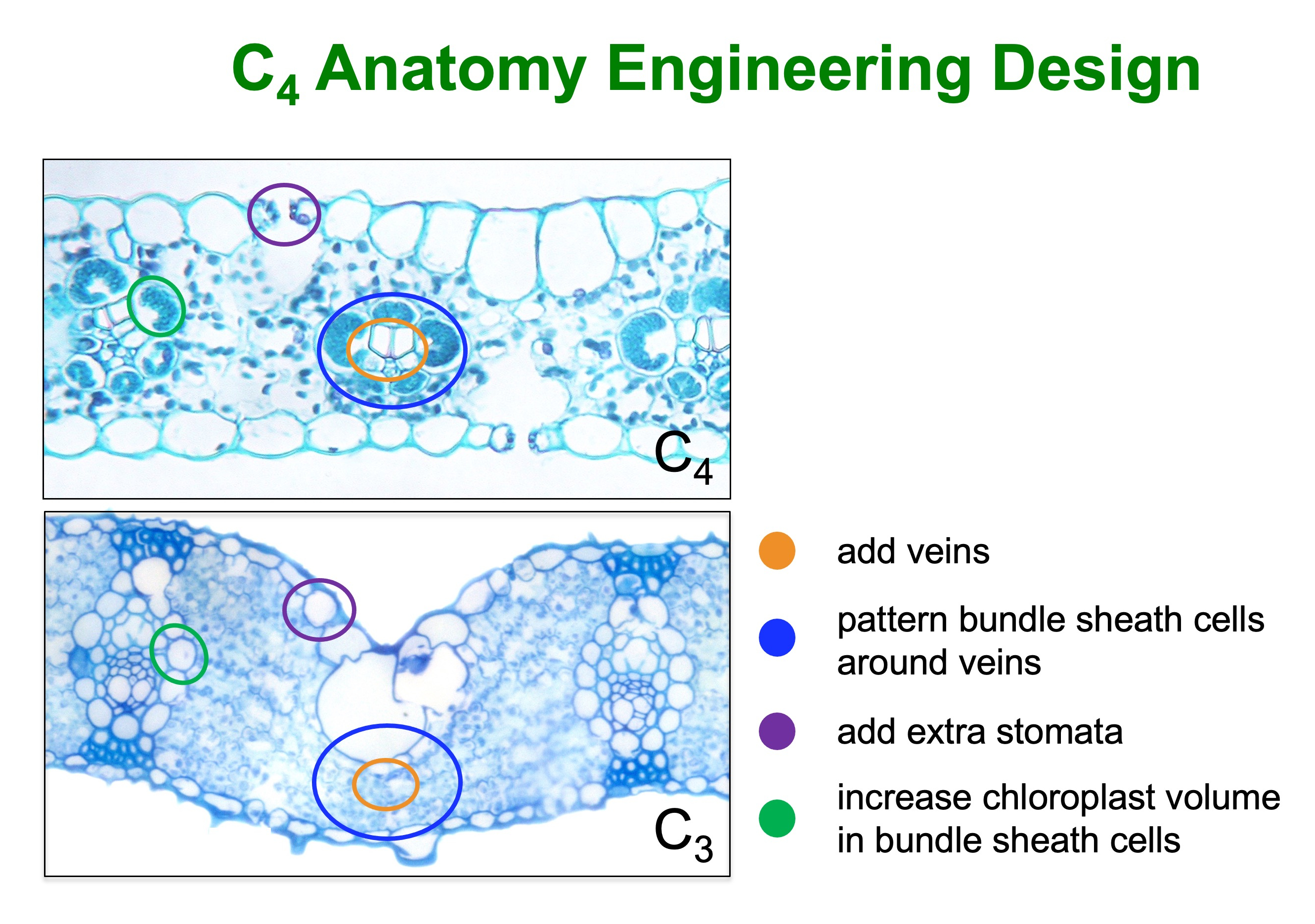
To introduce C4 ‘Kranz’ anatomy into rice, we need to change vein spacing patterns so that veins are closer together in the leaf, ensure bundle sheath cells are patterned around the extra veins, add stomatal rows in the epidermis above the extra veins and activate chloroplast development in the bundle sheath cells (rice bundle sheath cells are non-photosynthetic) (Figure 1). Given that it is not known how the development of Kranz anatomy is regulated in a C4 plant, much of our work is aimed at identifying regulatory genes in maize. However, engineering work in rice is also underway.
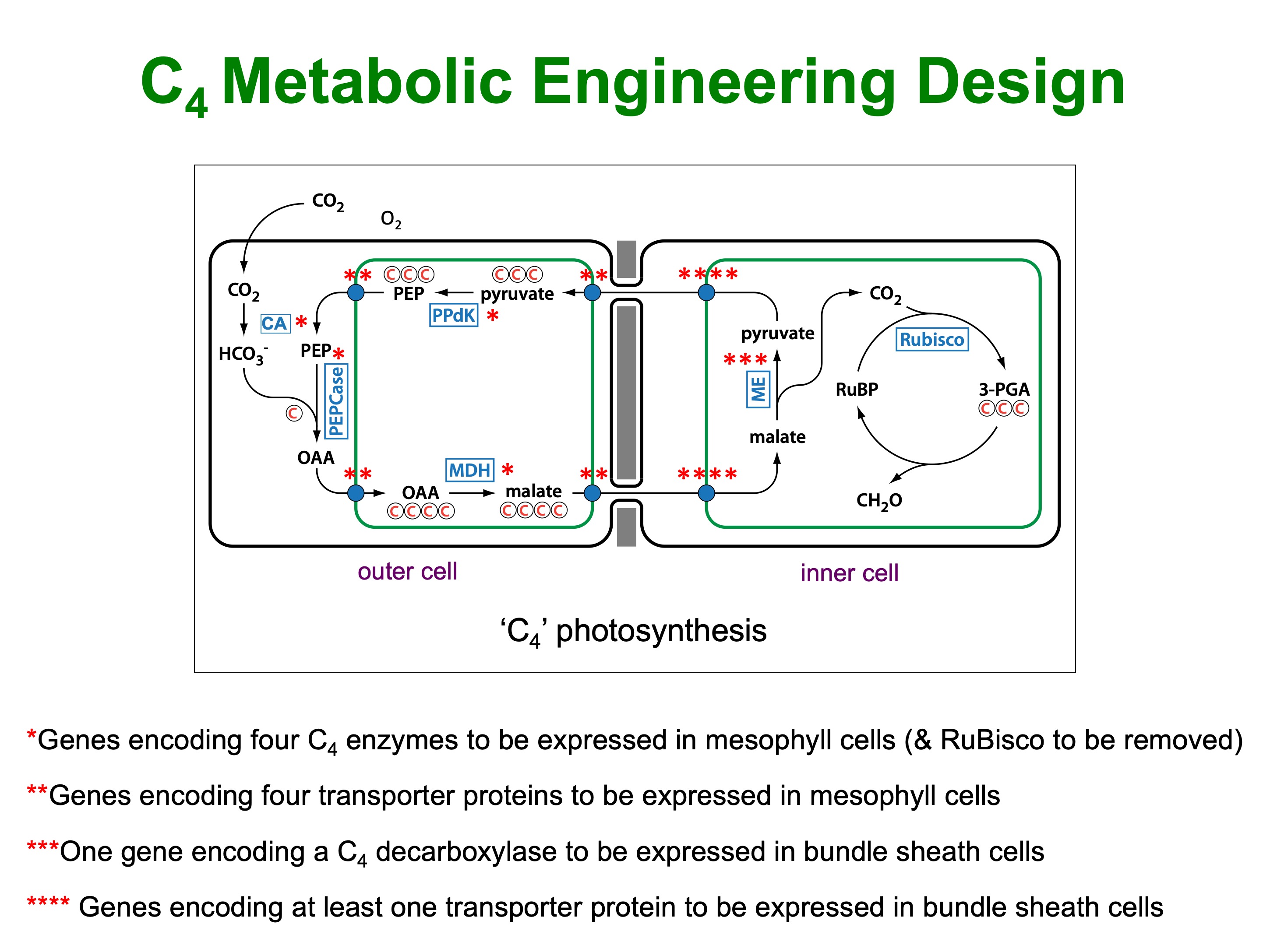
The route towards the introduction of C4 biochemistry is seemingly more straightforward, in that the genes encoding the C4 pathway enzymes and most of the metabolite transporters have been identified. However, there are at least 12 genes involved and they all need to be switched on at the right time, to the right level, and in the correct cell-type (Figure 2). For now, we are introducing the biochemistry into rice with C3 anatomy, with the goal of having a C4 cycle in a two cell radius of existing veins.
Ultimately, the two workstreams will come together and the biochemistry will be introduced into a rice leaf with Kranz anatomy. Some might say this is an unrealistic goal, so why do we think we can do it? Essentially we are being guided by evolution – the C4 pathway has evolved from the C3 pathway over 60 times independently and therefore, although the changes are quite complex, the transition could be relatively simple. We just need to work out the underlying mechanism.